As a bioengineer, Michael Kellen knows how to bridge worlds. It’s a skill that’s useful in his current role at Seattle-based Sage Bionetworks, where he works to import the technology and culture of the open-source software community into the medical research community.
Sage Bionetworks is a non-profit working to create open systems to hasten the pace of biomedical research. As director of technology platform and services, Kellen leads two teams: one that enables large-scale data sharing among life sciences researchers, and another that devises new ways to capture medical research data using cell phone and sensor platforms.
A foundation in software
As a bioengineering graduate student at UW in the lab of Professor James Bassingthwaighte, Kellen developed physiological modeling and simulation software that is still used in the open source community. Seattle-based software company Teranode later licensed and commercialized a version of his technology. After earning his Ph.D. in bioengineering in 2002 with a focus in computational biology, Kellen joined Teranode as the company’s first employee. He held positions in product development, field consulting, and development management until 2010, when he joined Sage Bionetworks.
New ways to collaborate on big data
Achieving an in-depth understanding of human diseases requires many different types of data from many sources, and a large number of clinical samples. “The notion that one small group or even one large company can lock up an area, understand it completely and get a product out is increasingly breaking down in life sciences and in the pharmaceutical industry, in which I interact a lot,” Kellen says.
Sage Bionetworks’ approach is to facilitate partnerships across institutions. Sage helps researchers join forces to tackle a large project and complex datasets. It also builds the general strategies and tools that can be used by many groups to solve diverse problems.
Building an open source culture
To encourage groups of people to choose to work together in a large-scale way, rather than work in their own individual lab groups, Kellen and his team developed Synapse, a platform that supports open, large-scale analysis of clinical genomics data. The technology may be powerful, he says, but first you need “the will to work together.”
Motivating people to want a collaboration to succeed in the first place and building trust is the greatest challenge, Kellen says. This can be even more difficult with complex technology and science problems. “Once they turn into technical challenges and not people challenges, it’s kind of inevitable that people find ways to make it work.”
Kellen says the vision for Synapse is a system in which scientists collaboratively build predictive models of human disease using large-scale, pooled data. Ultimately, these models could help evaluate a patient with a complicated disease such as cancer, Parkinson’s or Alzheimer’s –diseases that currently have huge variations in side effects and treatment effectiveness in different people –and then determine the best treatment options for an individual.
“I think the bottleneck is understanding the data, not generating the data,” Kellen says. “The measurement technologies are getting really good and really cheap amazingly quickly right now. The ability to use that data is lagging behind.”
A case study: colon cancer researchers pool resources
Over the last few years a number of groups conducted clinical and genomic studies on different patients with colon cancer. A string of papers from well-known groups followed, each explaining how they analyzed their data and found major sub-classifications of the disease, which can help guide treatment decisions. The first four teams published their articles, but none of them agreed on how the disease was breaking down into major sub-areas. By the time the fifth and sixth research teams published, there was massive confusion in the field about how the data was analyzed and what it meant, Kellen says.
Sage offered their Synapse technology platform to the group leaders, allowing them to pool their data. “We were able to get them to agree that if they each tried to do their own analysis on the pooled data, then they all had a reason to share their data with each other first and then eventually put that into the public domain and let other people work on it as well,” Kellen says. He expects the consortium of groups will release a paper soon that consolidates the independent studies and provides more consensus on the sub-typing classification system for the disease.
DREAM challenges, the ‘XPrizes’for bioinformatics
The Synapse platform also supports Sage’s DREAM Challenges, which are open competitions to crowd-source solutions to specific bioscience problems. In five recent competitions that finished in September 2014, more than 1,700 people signed up to compete. Challenge problems included identifying the best biomarkers for early Alzheimer’s-related cognitive decline and predicting how acute myeloid leukemia patients will fare after treatment. Four new challenges are currently underway – including projects to predict how a molecule will smell and forecast survival for prostate cancer patients. Prizes include invitations to speak at international conferences, publication in prestigious journals and high-profile publicity for the winners.
“It’s really interesting what happens when you don’t go to the five experts in an area and see what they think – you just leave it wide open,” Kellen says. “It is another aspect of leveraging data and the power of open communities of people in order to find solutions to problems.”
A bridge to social and sensor technology

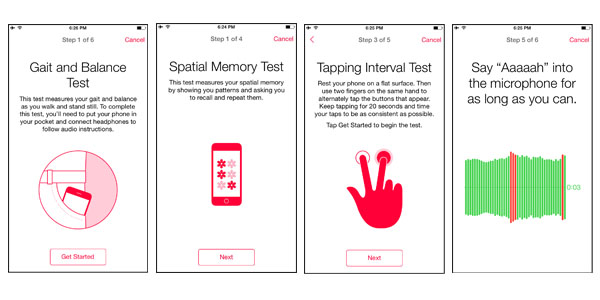
A Sage Bionetworks app enables patients with Parkinson’s disease to partner in research via Apple’s ResearchKit.
Kellen also leads a new area of research for Sage — mobile and sensor technologies that measure physiological aspects and traits of people living with disease. On March 9, Apple announced a new initiative called ResearchKit, an effort to jump-start medical research studies delivered through an iPhone. In partnership with Apple, Kellen’s technology team at Sage further developed their common back-end data capture service, called Bridge Server, for the iPhone. The service aggregates iPhone data collected from participating patients in real time and creates data sets that are distributed to researchers through Synapse. Sage’s service supports medical research apps in Parkinson’s disease and breast cancer developed by others at Sage, as well as apps in cardiovascular disease, diabetes and asthma developed by researchers at Stanford, Massachusetts General Hospital and Mount Sinai’s school of medicine.

“People are sharing more information on social networks and in gaming systems, so we thought, how can you make participating in a health study a little bit more engaging,” Kellen says. “With their smartphone, people can contribute their data to a research study and be a part of something that’s helping to create a better understanding of their disease.”
Solid core skills from UW BioE
Kellen says the UW BioE Ph.D. program taught him how to adapt to emerging fields and how to apply himself in a field of rapid changes. “The program really trains you to be able to think critically about a problem and be self-reliant, and to get original research out that is inherently multi-disciplinary and requires you to teach yourself a lot of different things,” he says. “Now that I’m more than 10 years out of the Ph.D. program, I appreciate the way I learned to think about the world and know that I have the ability to learn a new area, ramp up and then make progress. The training that I see going on in the bioengineering department is really good in that regard. It’s a very diverse set of areas that students are exposed to, and I think that serves them well in a variety of careers down the road.”
An industry pro recommends two key skill sets
Kellen advises current undergraduate and graduate-level bioengineering students to emphasize computer science and programming skills, regardless of their chosen field. “The ability to write your own code to get a problem solved is becoming one of the things that everybody needs to be able to do a little bit if they’re going into an engineering field,” he says. “I would also emphasize statistics and data science a bit more. That’s also one of those really transferrable skill sets where people will be working in a lot of different sub-disciplines and the specifics of what they’re studying are going to be very different, but there are some common statistical issues involved in working with data regardless of where it’s coming from.”
Some may recognize Kellen as a guest speaker at past UW BioE events. “Linking to the Bioengineering department is a good way to link to a variety of things that are going on in life sciences that are diverse and hard to follow,” Kellen says. “I always enjoy coming back to the department and getting a sense of all the different things that people are working on that I don’t encounter in my daily work.”
Article by Lia Unrau