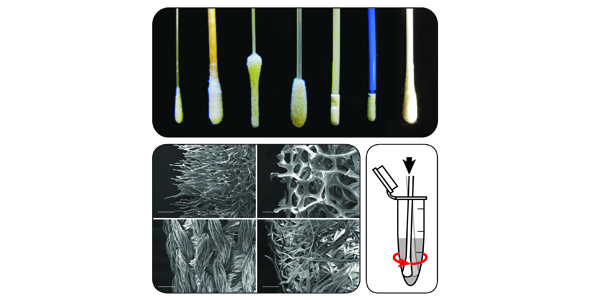
Routine diagnostic tests for a variety of different diseases share a common element, a swab used to capture a sample of a patient’s saliva, mucus or other bodily fluid and transfer the sample to a medium that can be used to run the test. However, not all swabs are created alike and differences between swab types may significantly affect a diagnostic test’s performance, negatively impacting its results.
UW Bioengineering Ph.D. student Nuttada Panpradist examined this clinical problem from an in-depth engineering perspective in a paper recently published in PLOS One. The paper presents experiments designed to quantitatively analyze how multiple types of swabs released specimen samples under various conditions relevant to rapid on-site testing. The results showed that commonly-used swab types varied drastically in their ability to release specimens, a complexity compounded by whether the sampling site was wet or dry.
Previously published work on swabs used in diagnostic testing focused on qualitative analysis or on clinical outcomes for specific diseases. Nuttada’s work establishes “some of the first objective, quantitative analysis of swab performance, which can be critically important when health care providers want to test patients for infectious disease,” explained collaborator and co-author Dr. Janet A. Englund, M.D., professor and clinician of pediatric infectious disease at UW and Seattle Children’s Hospital. “The results of this paper have implications for future development of point-of-care testing for important infections in developing countries as well as here at home,” she said.
The study’s results may inform future diagnostic test development, helping test developers select appropriate swab types and transfer methods for diagnosis of a wide variety of disease.
Nuttada has already begun to explore applications of this study’s findings. Her Ph.D. advisor UW Bioengineering Assistant Professor Dr. Barry Lutz said that Nuttada’s experience with sampling led her to develop an innovative swab sampling method for TB diagnostics from urine samples. This project recently received a Global WACh/W.H. Coulter Foundation Seed Grant.
The study was conducted as part of a large collaboration led by UW Bioengineering Professor Dr. Paul Yager to develop next-generation diagnostics that can be used in clinical environments, at home and in low-resource settings. The multiplexable autonomous disposable nucleic acid amplification test, or MAD NAAT, will be a low-cost, instrument-free test that detects nucleic acids (DNA or RNA) from multiple pathogens. Early targets include MRSA (methicillin-resistant staphylococcus aureus) and RSV (respiratory syncytial virus), commonly sampled from the nose.
Funding Statement:
The work presented in this paper was carried out in part with support from the Defense Advanced Research Projects Agency Defense Sciences Office under a grant to the University of Washington (grant number HR0011-11-2-0007, PI Paul Yager) with collaborators at PATH, GE Global Research, and ElitechGroup North America Inc./Epoch Biosciences. The funders had no role in study design, data collection and analysis, decision to publish, or preparation of the manuscript.